Abstract
The prevalence of comorbidities in individuals with neurodevelopmental disorders (NDDs) is not well understood, yet these are important for accurate diagnosis and prognosis in routine care and for characterizing the clinical spectrum of NDD syndromes. We thus developed PhenomAD-NDD, an aggregated database containing the comorbid phenotypic data of 51,227 individuals with NDD, all harmonized into Human Phenotype Ontology (HPO), with in total 3,054 unique HPO terms. We demonstrate that almost all congenital anomalies are more prevalent in the NDD population than in the general population, and the NDD baseline prevalence allows for an approximation of the enrichment of symptoms. For example, such analyses of 33 genetic NDDs show that 32% of enriched phenotypes are currently not reported in the clinical synopsis in the Online Mendelian Inheritance in Man (OMIM). PhenomAD-NDD is open to all via a visualization online tool and allows us to determine the enrichment of symptoms in NDD.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
The contents of PhenomAD-NDD are freely available online at https://humandiseasegenes.nl/phenomadndd/, as well as a tool to calculate the enrichment of symptoms for your own dataset. The summarized data are available at https://github.com/ldingemans/PhenomAD_NDD_data/. PubMed (https://pubmed.ncbi.nlm.nih.gov/) was used for the literature search in this study.
Code availability
If the tools provided online are insufficient, or if others would like to implement the enrichment analyses themselves, all code and summarized data used and developed in this study are available at https://github.com/ldingemans/PhenomAD_NDD_data/ (ref. 43).
References
Penrose, L. S. et al. A Clinical and Genetic Study of 1280 Cases of Mental Defect (HMSO, 1938).
Sarason, S. B. Psychological Problems in Mental Deficiency (Harper, 1949).
Yannet, H. Diagnostic classification of patients with mental deficiency: distribution of 1,330 institutionalized patients, with review of incidence of convulsive disorders and noncerebral developmental anomalies. Am. J. Dis. Child. 70, 83–88 (1945).
Kallmann, F. J. Heredity in Health and Mental Disorder; Principles of Psychiatric Genetics in the Light of Comparative Twin Studies (Norton, 1953).
Pitt, D. & Roboz, P. A survey of 782 cases of mental deficiency. J. Ment. Defic. Res. 9, 4–23 (1965).
Gustavson, K. H., Holmgren, G., Jonsell, R. & Son Blomquist, H. K. Severe mental retardation in children in a northern Swedish county. J. Ment. Defic. Res. 21, 161–180 (1977).
Laxova, R., Ridler, M. A. & Bowen-Bravery, M. An etiological survey of the severely retarded Hertfordshire children who were born between January 1, 1965 and December 31, 1967. Am. J. Med. Genet. 1, 75–86 (1977).
Göstason, R. Psychiatric illness among the mentally retarded. A Swedish population study. Acta Psychiatr. Scand. Suppl. 318, 1–117 (1985).
Reid, A. H. Schizophrenia in mental retardation: clinical features. Res. Dev. Disabil. 10, 241–249 (1989).
Deb, S. & Joyce, J. Psychiatric illness and behavioural problems in adults with learning disability and epilepsy. Behav. Neurol. 11, 125–129 (1999).
Holland, T. & Murphy, G. Behavioural and psychiatric disorder in adults with mild learning difficulties. Int. Rev. Psychiatry 2, 117–135 (1990).
Mirabi, M. in Child and Adolescent Psychiatry, Mental Retardation and Geriatric Psychiatry (eds Pichot, P. et al.) 177–181 (Springer, 1985).
Corbett, J. A. in Scientific Studies in Mental Retardation (eds Dobbing, J. et al.) 479–506 (Palgrave Macmillan, 1984).
Linna, S. L. et al. Psychiatric symptoms in children with intellectual disability. Eur. Child Adolesc. Psychiatry 8, 77–82 (1999).
Farag, T. I. et al. Disease profile of 400 institutionalized mentally retarded patients in Kuwait. Clin. Genet. 44, 329–334 (1993).
Bratek, A., Krysta, K. & Kucia, K. Psychiatric comorbidity in older adults with intellectual disability. Psychiatr. Danub. 29, 590–593 (2017).
McGrother, C. W. et al. Epilepsy in adults with intellectual disabilities: prevalence, associations and service implications. Seizure 15, 376–386 (2006).
Daily, D. K., Ardinger, H. H. & Holmes, G. E. Identification and evaluation of mental retardation. Am. Fam. Physician 61, 1059–1067 (2000).
Karczewski, K. J. et al. The mutational constraint spectrum quantified from variation in 141,456 humans. Nature 581, 434–443 (2020).
Byrd, J. B., Greene, A. C., Prasad, D. V., Jiang, X. & Greene, C. S. Responsible, practical genomic data sharing that accelerates research. Nat. Rev. Genet. 21, 615–629 (2020).
Robinson, P. N. et al. The Human Phenotype Ontology: a tool for annotating and analyzing human hereditary disease. Am. J. Hum. Genet. 83, 610–615 (2008).
Köhler, S. et al. The human phenotype ontology in 2021. Nucleic Acids Res. 49, D1207–D1217 (2021).
Firth, H. V. et al. DECIPHER: Database of Chromosomal Imbalance and Phenotype in Humans using Ensembl Resources. Am. J. Hum. Genet. 84, 524–533 (2009).
Boyd, P. A. et al. Paper 1: the EUROCAT network—organization and processes. Birth Defects Res. A 91, S2–S15 (2011).
Dolk, H. EUROCAT: 25 years of European surveillance of congenital anomalies. Arch. Dis. Child. Fetal Neonatal Ed. 90, F355–F358 (2005).
Greenlees, R. et al. Paper 6: EUROCAT member registries: organization and activities. Birth Defects Res. A 91, S51–S100 (2011).
McKusick, V. A. Mendelian Inheritance in Man and its online version, OMIM. Am. J. Hum. Genet. 80, 588–604 (2007).
Nabais Sá, M. J. et al. De novo and biallelic DEAF1 variants cause a phenotypic spectrum. Genet. Med. 21, 2059–2069 (2019).
Fokkema, I. F. A. C. et al. LOVD v.2.0: the next generation in gene variant databases. Hum. Mutat. 32, 557–563 (2011).
Adler, A. et al. PhenoDis: a comprehensive database for phenotypic characterization of rare cardiac diseases. Orphanet J. Rare Dis. 13, 22 (2018).
Li, B. et al. GPCards: an integrated database of genotype-phenotype correlations in human genetic diseases. Comput. Struct. Biotechnol. J. 19, 1603–1611 (2021).
Mungall, C. J. et al. The Monarch Initiative: an integrative data and analytic platform connecting phenotypes to genotypes across species. Nucleic Acids Res. 45, D712–D722 (2017).
Lek, M. et al. Analysis of protein-coding genetic variation in 60,706 humans. Nature 536, 285–291 (2016).
Fuller, Z. L., Berg, J. J., Mostafavi, H., Sella, G. & Przeworski, M. Measuring intolerance to mutation in human genetics. Nat. Genet. 51, 772–776 (2019).
Lelieveld, S. H. et al. Meta-analysis of 2,104 trios provides support for 10 new genes for intellectual disability. Nat. Neurosci. 19, 1194–1196 (2016).
Kaplanis, J. et al. Evidence for 28 genetic disorders discovered by combining healthcare and research data. Nature 586, 757–762 (2020).
Dingemans, A. J. M. et al. Human disease genes website series: an international, open and dynamic library for up-to-date clinical information. Am. J. Med. Genet. A 185, 1039–1046 (2021).
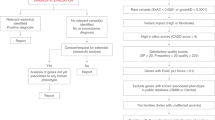
Page, M. J. et al. The PRISMA 2020 statement: an updated guideline for reporting systematic reviews. Int. J. Surg. 88, 105906 (2021).
Bokeh Development Team. Bokeh: Python Library for Interactive Visualization https://bokeh.pydata.org/en/latest/ (2021).
Manders, P., Lutomski, J. E., Smit, C., Swinkels, D. W. & Zielhuis, G. A. The Radboud Biobank: a central facility for disease-based biobanks to optimise use and distribution of biomaterial for scientific research in the Radboud University Medical Center, Nijmegen. Open J. Bioresour. 5, 2 (2018).
Richards, S. et al. Standards and guidelines for the interpretation of sequence variants: a joint consensus recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genet. Med. 17, 405–424 (2015).
Daoud, J. Multicollinearity and regression analysis. J. Phys. Conf. Ser. 949, 012009 (2017).
Dingemans, L. ldingemans/PhenomAD_NDD_data: original release as in the paper. Zenodo https://doi.org/10.5281/zenodo.10908427 (2024).
Acknowledgements
We are grateful to the Dutch Organisation for Health Research and Development for ZON-MW grant no. 912-12-109 (to B.B.A.d.V. and L.E.L.M.V.), Donders Junior researcher grant no. 2019 (to B.B.A.d.V. and L.E.L.M.V.) and Aspasia grant no. 015.014.066 (to L.E.L.M.V.). The aims of this study contribute to the Solve-RD project (to L.E.L.M.V.), which has received funding from the European Union’s Horizon 2020 Research and Innovation program under grant agreement no. 779257. Multiple authors of this publication are members of the European Reference Network on Rare Congenital Malformations and Rare Intellectual Disability ERN-ITHACA (EU Framework Partnership Agreement ID: 3HP-HPFPA ERN-01-2016/739516). The funder(s) of the study had no role in study design, data collection, data analysis, data interpretation or writing of the report.
Author information
Authors and Affiliations
Contributions
This research included worldwide researchers who were involved throughout the research process and each had their own role and responsibility. The study was designed by L.E.L.M.V. and B.B.A.d.V., and implemented by A.J.M.D., L.E.L.M.V. and B.B.A.d.V. The data are owned by the various independent researchers (A.J.M.D., S.J., J.v.R., N.d.L., R.P., J.S.-H., B.W.v.B., C.M., C.W.O., M.W., P.J.v.d.S., G.W.E.S., R.F.K., A.T.V.-v.S., T.K., D.A.K., L.E.L.M.V. and B.B.A.d.V.), whereas the intellectual property lies with the leading research group from the Radboud University Medical Center. The original draft was written by A.J.M.D., L.E.L.M.V. and B.B.A.d.V. All authors were involved in reviewing and editing the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Medicine thanks Kristopher Kahle and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editor: Sonia Muliyil, in collaboration with the Nature Medicine team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Countries contributing to EUROCAT.
The 18 countries whose institutions contribute data to the EUROCAT registry, with their relative contribution in various shades of blue. Of note, the total number of individuals in the EUROCAT registry is 11,616,332.
Extended Data Fig. 2 Distribution of number of clinical features.
The distribution of the HPO terms of the individuals seen at the outpatient clinic in the Radboud University Medical Center.
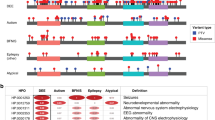
Extended Data Fig. 3 Result of enrichment analysis.
A) The results of the top 10 results of the enrichment analysis for two of the 33 investigated genetic syndromes are shown here: MRD50 (OMIM #617787), for which 16% of the symptoms are described in the clinical synopsis in OMIM, and NEDHELS (OMIM #617171, 100% concordance). Shown are the 10 clinical features that are the most enriched: that is with the highest ratio when dividing the prevalence in the syndrome with the prevalence in PhenomAD-NDD. The size of the nodes in the network corresponds to the prevalence in that specific genetic syndrome. The phenotypic data of the investigated syndromes was gathered from the specific website of the Human Disease Gene website series, with the prevalence rate in the syndrome referring to the proportion of individuals with the specific genetic syndrome who exhibit a particular clinical feature. The color corresponds to the enrichment ratio, which compares the prevalence in the syndrome to the prevalence in PhenomAD-NDD. To avoid spurious enrichments, only clinical features with a prevalence of at least 1% are shown in this figure. Using the developed tool in either the electronic supplement or online at the Human Disease Genes website series (https://humandiseasegenes.nl/phenomadndd/)6, others are able to determine enriched clinical features and generate these figures for their syndrome of interest. B) The clinical synopsis of OMIM8 as shown for these two genetic syndromes.
Supplementary information
Supplementary Table 1
The search strategy of the literature search and meta-analysis, including all included and excluded studies and data collected per study.
Supplementary Table 2
The results of the enrichment analysis for 33 genetic disorders.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Dingemans, A.J.M., Jansen, S., van Reeuwijk, J. et al. Prevalence of comorbidities in individuals with neurodevelopmental disorders from the aggregated phenomics data of 51,227 pediatric individuals. Nat Med 30, 1994–2003 (2024). https://doi.org/10.1038/s41591-024-03005-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41591-024-03005-7